Normative Connectomes
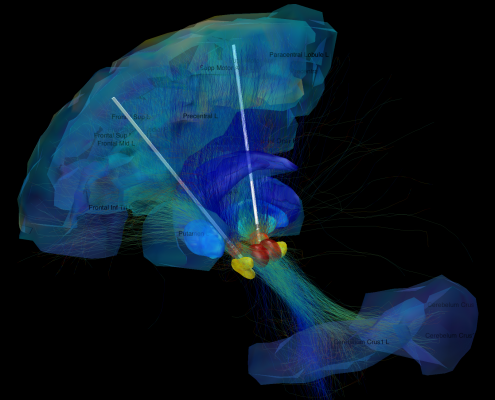
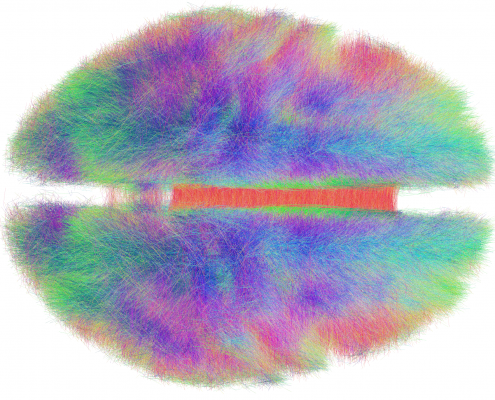
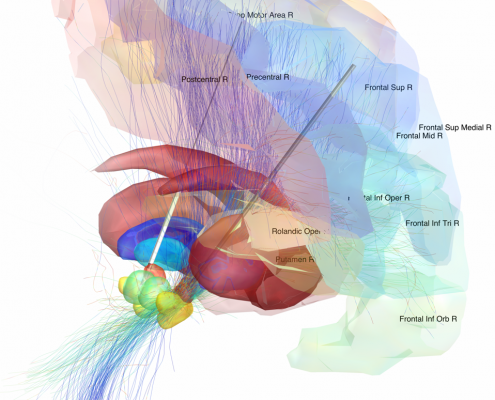
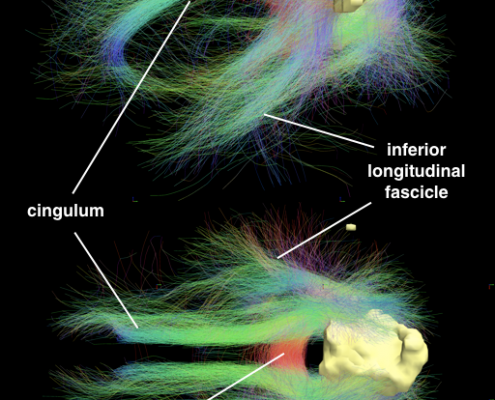
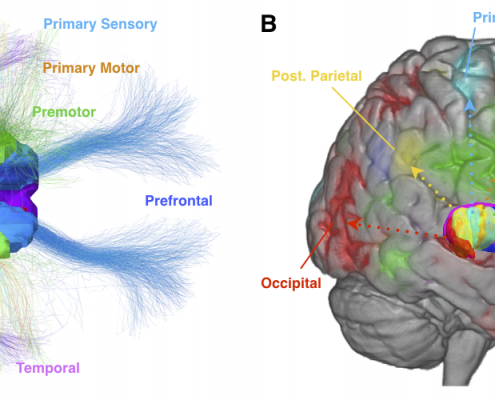
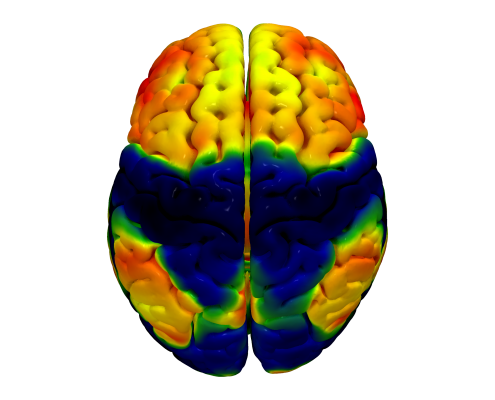
Since 2014, we worked on estimating “average wiring diagrams” of the human brain. Based on diffusion-weighted or resting-state functional MRI data of large cohorts of subjects/patients, this line of work focused on running whole-brain tractography or functional connectivity analyses, warping individual connectomes into standard space and then averaging the results across the group of subjects.
In 2014, we published a first article under the lead of Felix Blankenburg describing the effort of a first structural-functional MNI connectome. There is a small additional article associated with the data release of this first connectome. 2016, we replicated results based on a much larger cohort of healthy subjects and validated the use of such normative connectomes by replicating multiple connectomic studies directly within template space and based on the group connectome only.
In 2017, a first article under the lead of Andrea A Kühn made use of such normative connectomes by combining connectivity measures with electrophysiological recordings of the subcortex. This novel connectome in use here was based on high-quality multishell diffusion data acquired within the human connectome project (Setsompop 2013).
A second article published under the lead of Michael D Fox explored connectivity profiles of literature based deep brain stimulation targets using normative connectomes. This made use of a 1000-healthy subject functional connectome based on data off the Harvard genomic superstruct project that is also used on neurosynth.org and was both preprocessed and provided by Thomas Yeo (Yeo 2011).
Finally, within work of a third article, both functional and structural average connectomes from patients suffering from Parkinson’s Disease were created (using data of the PPMI project; ppmi-info.org).
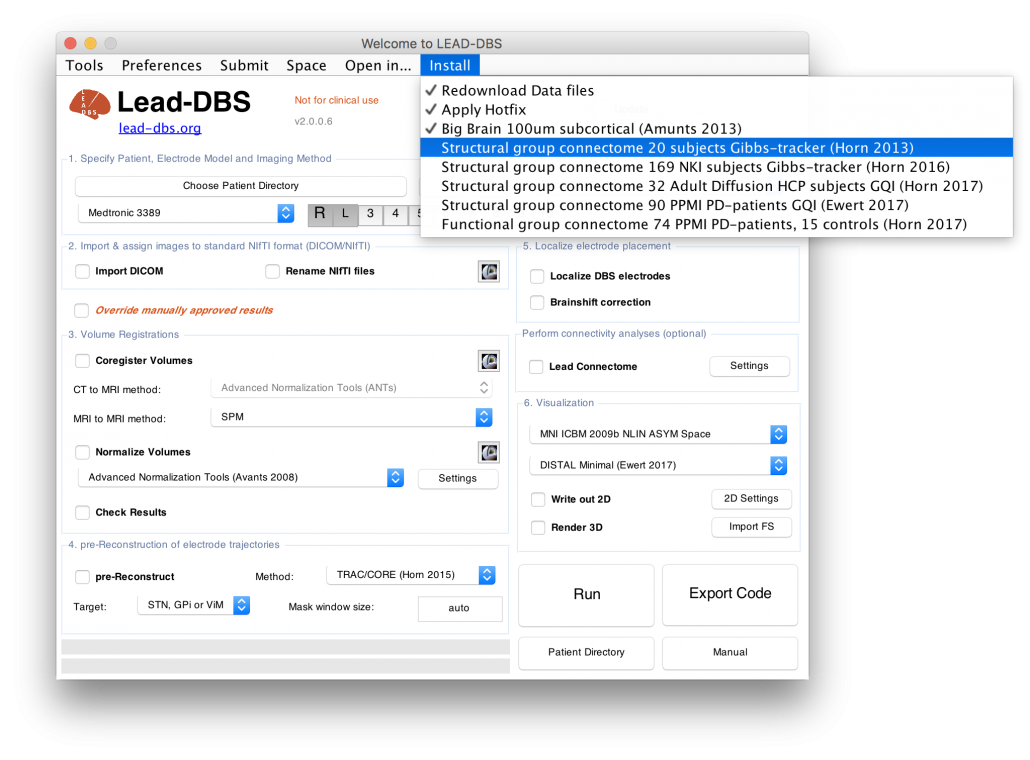
Most normative connectomes are available for direct download within the Lead Neuroimaging Suite (leadsuite.io)
List of currently available connectomes
- Structural normative connectomes
- Structural group connectome 19 subjects Gibbs-tracker (Horn 2013)
[Direct download current version v.1.0] - Structural group connectome 169 NKI-RS subjects Gibbs-tracker (Horn 2016)
[Direct download current version v.1.0; Download via NITRC] - Structural group connectome 32 Adult Diffusion MGH-USC HCP subjects GQI (Dataset: Setsompop 2013; processing: Horn 2017)
[Direct download current version v.1.2; older versions: v.1.1, v.1.0] - Structural group connectome 85 PPMI PD-patients GQI (Dataset: Marek 2011; processing: Ewert 2017)
[Direct download current version v.1.1; older versions: v.1.0]
- Structural group connectome 19 subjects Gibbs-tracker (Horn 2013)
- Functional normative connectomes
- Functional group connectome 1000 GSP healthy subjects, can be directly used for network mapping within Lead-DBS (Lead Mapper and NetworkMapping Explorer), preprocessed by Bassam Al-Fatly. [Download from Harvard Dataverse]
- Functional group connectome 74 PPMI PD-patients, 15 controls (Dataset: Marek 2011; processing: Horn 2017)
[Direct download current version v.1.0] - Functional group connectome 75 Toronto PD-patients (Detailed description: Zenodo; processing: Horn 2017)
[Direct download current version v1.0]
Applications
Normative connectomes cannot replace patient/subject-specific connectivity information but are useful in scenarios where subject-specific data is not available and cannot be acquired. This often includes clinical applications such as:
- Deep Brain Stimulation: After surgery, patients cannot be scanned in the MRI without restrictions, acquiring high quality data is difficult if not impossible (e.g. Horn 2017a,b,c).
- Other forms of Brain Stimulation: Similar concepts may apply to noninvasive brain stimulation such as TMS (e.g. Weigand 2017)
- Stroke: After stroke, patients can undergo MRI scans, but their brain structure is modified by the stroke event. If researchers are interested in how connectivity was before the stroke event, normative connectomes may at least give a useful hint (e.g. Boes 2015, Laganiere 2016, Fischer 2016, Darby 2016 & 2017, Fasano 2017)
- Analysing large databases: It is often difficult if not impossible to acquire connectivity data from large cohorts of patients/subjects that are analysed retrospectively. Still, such databases can sometimes yield highly valuable information and the combination with normative connectomes can yield further information (e.g. Horn 2017a).